Note
Go to the end to download the full example code
ERP EEG decoding in Tangent space.¶
Decoding applied to EEG data in sensor space decomposed using Xdawn. After spatial filtering, covariances matrices are estimated, then projected in the tangent space and classified with a logistic regression.
# Authors: Alexandre Barachant <alexandre.barachant@gmail.com>
#
# License: BSD (3-clause)
import numpy as np
from matplotlib import pyplot as plt
import mne
from mne import io
from mne.datasets import sample
from sklearn.linear_model import LogisticRegression
from sklearn.model_selection import KFold
from sklearn.metrics import confusion_matrix, ConfusionMatrixDisplay
from sklearn.pipeline import make_pipeline
from pyriemann.estimation import XdawnCovariances
from pyriemann.tangentspace import TangentSpace
print(__doc__)
Set parameters and read data¶
data_path = str(sample.data_path())
raw_fname = data_path + "/MEG/sample/sample_audvis_filt-0-40_raw.fif"
event_fname = data_path + "/MEG/sample/sample_audvis_filt-0-40_raw-eve.fif"
tmin, tmax = -0.0, 1
event_id = dict(aud_l=1, aud_r=2, vis_l=3, vis_r=4)
# Setup for reading the raw data
raw = io.Raw(raw_fname, preload=True, verbose=False)
raw.filter(2, None, method="iir") # replace baselining with high-pass
events = mne.read_events(event_fname)
raw.info["bads"] = ["MEG 2443"] # set bad channels
picks = mne.pick_types(
raw.info, meg=False, eeg=True, stim=False, eog=False, exclude="bads"
)
# Read epochs
epochs = mne.Epochs(
raw,
events,
event_id,
tmin,
tmax,
proj=False,
picks=picks,
baseline=None,
preload=True,
verbose=False,
)
labels = epochs.events[:, -1]
evoked = epochs.average()
epochs_data = epochs.get_data(copy=False)
Filtering raw data in 1 contiguous segment
Setting up high-pass filter at 2 Hz
IIR filter parameters
---------------------
Butterworth highpass zero-phase (two-pass forward and reverse) non-causal filter:
- Filter order 8 (effective, after forward-backward)
- Cutoff at 2.00 Hz: -6.02 dB
Removing projector <Projection | PCA-v1, active : False, n_channels : 102>
Removing projector <Projection | PCA-v2, active : False, n_channels : 102>
Removing projector <Projection | PCA-v3, active : False, n_channels : 102>
Decoding in tangent space with a logistic regression¶
n_components = 2 # pick some components
# Define a monte-carlo cross-validation generator (reduce variance):
cv = KFold(n_splits=10, shuffle=True, random_state=42)
clf = make_pipeline(
XdawnCovariances(n_components),
TangentSpace(metric="riemann"),
LogisticRegression(),
)
preds = np.zeros(len(labels))
for train_idx, test_idx in cv.split(epochs_data):
y_train, y_test = labels[train_idx], labels[test_idx]
clf.fit(epochs_data[train_idx], y_train)
preds[test_idx] = clf.predict(epochs_data[test_idx])
# Printing the results
acc = np.mean(preds == labels)
print("Classification accuracy: %f " % (acc))
names = ["audio left", "audio right", "vis left", "vis right"]
cm = confusion_matrix(labels, preds)
ConfusionMatrixDisplay(cm, display_labels=names).plot()
plt.show()
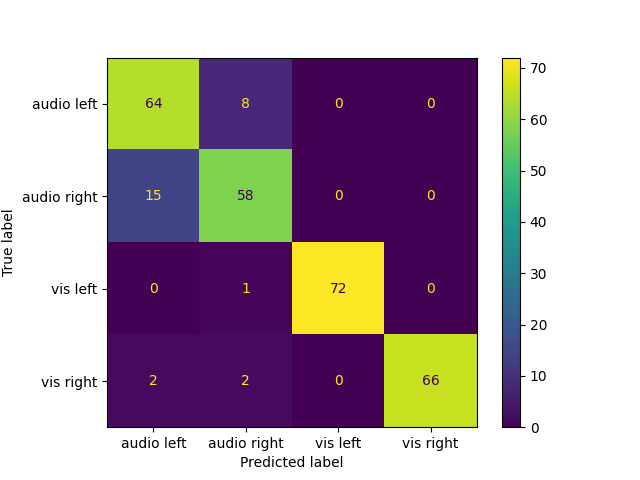
Classification accuracy: 0.902778
Total running time of the script: (0 minutes 5.301 seconds)