pyRiemann: Biosignals classification with Riemannian geometry¶
pyRiemann is a Python machine learning package based on scikit-learn API. It provides a high-level interface for processing and classification of real (resp. complex)-valued multivariate data through the Riemannian geometry of symmetric (resp. Hermitian) positive definite (SPD) (resp. HPD) matrices.
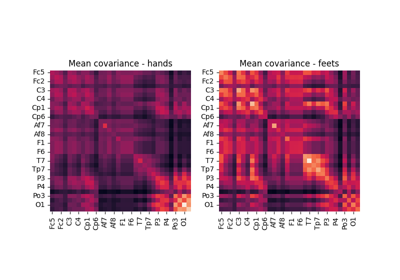
pyRiemann aims at being a generic package for multivariate data analysis but has been designed around biosignals (like EEG, MEG or EMG) manipulation applied to brain-computer interface (BCI), estimating covariance matrices from multichannel time series, and classifying them using the Riemannian geometry of SPD matrices.
For a brief introduction to the ideas behind the package, you can read the introductory notes. More practical information is on the installation page. You may also want to browse the example gallery to get a sense for what you can do with pyRiemann and API reference to find out how.
To see the code or report a bug, please visit the github repository.